Recording physiological signals during fMRI scans
TL;DR Attach the physio devices, switch on physio recording in the CMRR Multiband EPI Sequences Sequence/Special card by selecting DICOM from the drop-down menu, record your EPI data, and enjoy.
Introduction
Physiological signals such as heart rate and respiration can influence the fMRI BOLD signal (more below), so recording these signals can be useful for later processing and analysis. At the suggestion of a friendly expert, I explored how to record physiological signals at our institution's research MRI facility.
Happily, this proved to be pretty straightforward on our Siemens Prisma. However, it took me a while to find the necessary information on-line, hence this post.
Record
If you're using a recent build of the CMRR Multiband EPI Sequences, you're 99% of the way there. The hardworking CMRR experts have baked physiological recording into the sequence software itself; all you need to do is activate it. At the scanner console, browse to the Sequence/Special card and look to the top-right for a drop-down menu.
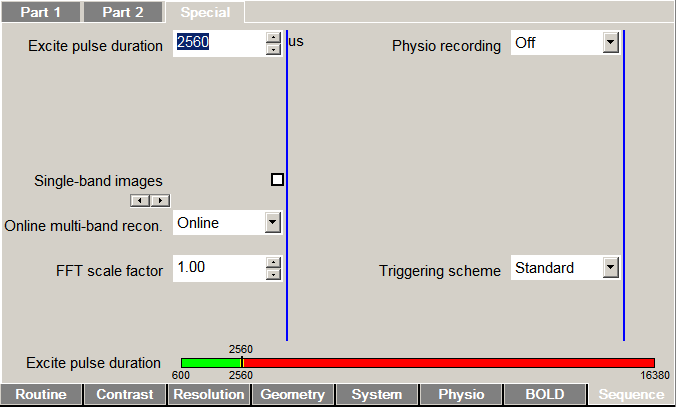
Lifting flagrantly from the sequence manual:
Physio. recording: Control recording of physiological signals (cardiac, respiration, ECG, external) to text or encoded DICOM files. Text files will be stored in C:\MedCom\Log\Physio\ on the host computer (MRC). Legacy uses the classic CPmuSequence/IdeaCmdTool logging facility (this is the only option available on VAxx/VBxx systems and VD11x; it is known to be unreliable on VD11x/VD13x/VE11x). DICOM and File use the new online/ICE logging available since VD13A, which is recommended for use when available. File writes the log data to individual text files for each signal (this option was named Online in previous versions). DICOM embeds these logs in a special DICOM “image” stored in the database (sample Matlab code for reading the special DICOM files is provided in the GitHub repository). Multiple enables both Legacy and File options (not recommended for routine use).
To test this, we put a volunteer in the scanner, hooked up the wireless respiratory and pulse-ox sensors, selected the DICOM option from drop-down menu on the Special card, and started recording EPI data. Lo and behold, we found a DICOM file associated with the study. Success!1
Extract
With the physio data saved to a DICOM, the next step was to extract the recordings. The CMRR repo has a Matlab script for this purpose, but I wanted to get a Python version working. This was down partly to Python chauvinism and partly to wanting a better understanding of what extraction is doing.
My draft implementation is available in a Github gist . The script takes the path to a physio DICOM as an argument (along with an optional output directory). The physiological recordings are then extracted to their original filenames in the output directory (same as the DICOM's if not supplied).
usage: physio_convert.py [-h] [--path PATH] [--outdir OUTDIR]
optional arguments:
-h, --help show this help message and exit
--path PATH path of physio DICOM file
--outdir OUTDIR directory for output
Depending on the data you collected at scan time, running the script should produce two or three of the following (the * is shared date-time and scan information):
| Signal | Filename | Sampling Rate (Hz) |
|---|---|---|
| Pulse | Physio_*_PULS.log |
200 |
| Respiration | Physio_*_RESP.log |
50 |
| EPI | Physio_*_Info.log |
Per parameters2 |
Inspect
The recordings themselves may require additional attention prior to use (this will depend what you want to do with them). As a first step, I wanted to visualize the data to ensure that we got something reasonable.
All signals are timestamped by clock "TICS", which on our scanner appear to be 400 Hz/2.5 msec timesteps. Although all signals are stamped with codes corresponding to this 400-Hz "heartbeat" signal, different signals are sampled and recorded at different rates. On our Prisma, we observed the sampling rates reported in the table above.
Aligning the signals is relatively painless using Python and Pandas. Plotting the data, we see the expected sinusoids for pulse (blue) and respiration (orange) as well as a steady pulse for EPI volumes (green). 😎
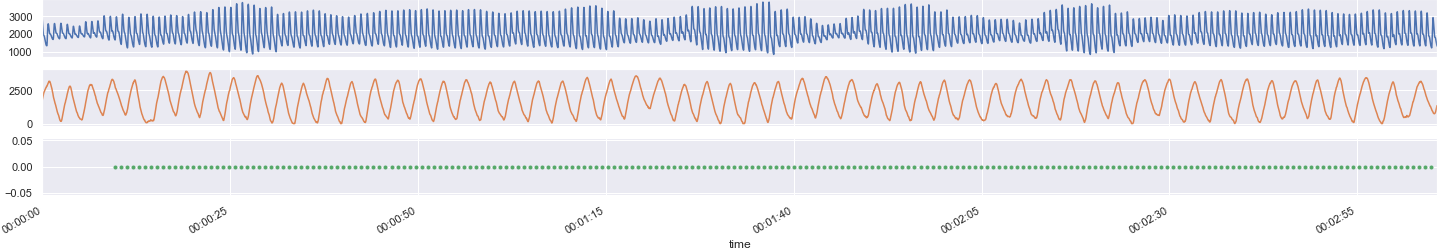
Regarding use of the physio recordings as covariates for fMRI processing and analysis, I'll defer to the expertise of other investigators. Some useful resources might include:
- Glover, G. H., Li, T. Q., & Ress, D. (2000). Image-based method for retrospective correction of physiological motion effects in fMRI: RETROICOR. Magnetic Resonance in Medicine, 44(1), 162–167.
- Birn, R. M., Diamond, J. B., Smith, M. A., & Bandettini, P. A. (2006). Separating respiratory-variation-related fluctuations from neuronal-activity-related fluctuations in fMRI. NeuroImage, 31(4), 1536–1548. https://doi.org/10.1016/j.neuroimage.2006.02.048
- Power, J. D., Lynch, C. J., Silver, B. M., Dubin, M. J., Martin, A., & Jones, R. M. (2019). Distinctions among real and apparent respiratory motions in human fMRI data. NeuroImage, 201, 116041. https://doi.org/10.1016/j.neuroimage.2019.116041
Acknowledgments
Thanks to the CMRR team for making this easy, to our scanner tech Lisa for her patience, and to Jonathan for reassurance that getting physio recordings from new scanners isn't too arduous.
The Aguirre lab has a resource page that provided useful hints.
Footnotes
1 Acquiring physio recording without the CMRR MB-EPI sequence is left as an exercise for the reader.
2 The timing information in the Info file includes start/stop for each volumes, slices, and (if used) multiple echoes. Having this information might be useful for various reasons including converting DICOM to NIFTI accurately, although tools such as dcm2niix should have the necessary timing information built-in.

